College of Engineering
Shining Light Makes Materials Magnetic at Room Temperature
In April, Engineering Professor Alexander Balatsky published his work applying AI to his theory of 'dynamic multiferroicity'
April 17, 2024 | Christie Wang, for the Office of the Vice President for Research
Engineering Entrepreneurship: Hacking For Defense
The course uses a project-based approach to get students out of the classroom and into the community, engaging with defense industry professionals
April 16, 2024 | Claire Tremont
UConn Leading Federally Backed Regional Initiative to Defend Electric Grid from Cyberattack
CyberCARED will take UConn’s cybersecurity research and development into a new domain
April 9, 2024 | Jaclyn Severance
UConn’s Engineers Without Borders Hosts Northeast Regional Conference
Students learned how other EWB chapters in the Northeast are helping communities around the world solve problems
April 3, 2024 | Olivia Drake
Graduate Students Share Research, Network with Peers at UConn’s Sustainability Summit
The inaugural summit, hosted by the College of Engineering's Center for Clean Energy Engineering (C2E2), brought together students from various disciplines within C2E2 and across at least five engineering departments and schools
April 2, 2024 | Jordan Baker
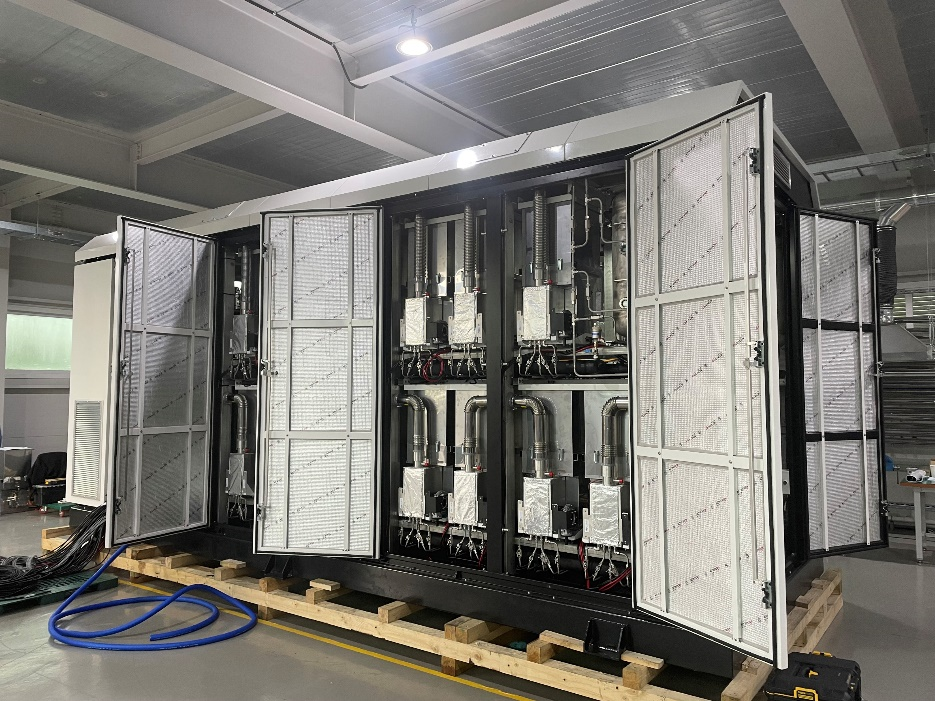
Gift of Fuel Cell Units Enhances UConn’s Clean Energy Commitment
Eight solid oxide fuel cell units will be donated to the Center for Clean Energy Engineering
April 2, 2024 | Matt Engelhardt
Professor Receives a $3M RO1 Grant From the National Institutes of Health
The research conducted at the UConn College of Engineering is poised to make significant contributions to the field of biomechanics and improve human health outcomes in the process.
March 29, 2024 | Joanna Giano
Engineering Entrepreneurship: UConn Students Participate in DOE Contest
These teams are working to develop attainable, equitable, scalable energy technologies and business opportunities.
March 21, 2024 | Claire Tremont
UConn Awarded $4.5M DOE Grant to Benefit Grid Reliability for Transmission and Distribution Systems
The project team will create open-source data visualization tools to display information about renewable energy sources and distributed energy resources
March 21, 2024 | Olivia Drake
UConn Successfully Pursues Energy Efficiency, Even As Campus Grows
Despite 1 million square feet of new building space, UConn's carbon footprint is smaller than it was 20 years ago
March 21, 2024 | Kim Krieger