College of Engineering
College of Engineering Names New Dean
Ji-Cheng ‘JC’ Zhao brings 30 years' experience working in academia, industry, and government
April 19, 2024 | Claire Tremont
Six UConn Faculty Members Named AAAS Fellows
The AAAS is the world’s largest general scientific society and publisher of the Science family of journals
April 18, 2024 | Mike Enright '88 (CLAS), University Communications
Meeting the High-Speed Challenge
The Air Force Research Laboratory awarded UConn an additional $10.5 million for projects related to welding and advanced materials for high-temperature applications
April 18, 2024 | Matt Engelhardt
UConn Celebrates Promotion and Tenure of 91 Faculty
Evaluations for promotion, tenure, and reappointment apply the highest standards of professional achievement in scholarship, teaching, and service for each faculty member evaluated
April 17, 2024 | Alexis Lohrey, Office of the Provost
Shining Light Makes Materials Magnetic at Room Temperature
In April, Engineering Professor Alexander Balatsky published his work applying AI to his theory of 'dynamic multiferroicity'
April 17, 2024 | Christie Wang, for the Office of the Vice President for Research
Engineering Entrepreneurship: Hacking For Defense
The course uses a project-based approach to get students out of the classroom and into the community, engaging with defense industry professionals
April 16, 2024 | Claire Tremont
UConn Leading Federally Backed Regional Initiative to Defend Electric Grid from Cyberattack
CyberCARED will take UConn’s cybersecurity research and development into a new domain
April 9, 2024 | Jaclyn Severance
UConn’s Engineers Without Borders Hosts Northeast Regional Conference
Students learned how other EWB chapters in the Northeast are helping communities around the world solve problems
April 3, 2024 | Olivia Drake
Graduate Students Share Research, Network with Peers at UConn’s Sustainability Summit
The inaugural summit, hosted by the College of Engineering's Center for Clean Energy Engineering (C2E2), brought together students from various disciplines within C2E2 and across at least five engineering departments and schools
April 2, 2024 | Jordan Baker
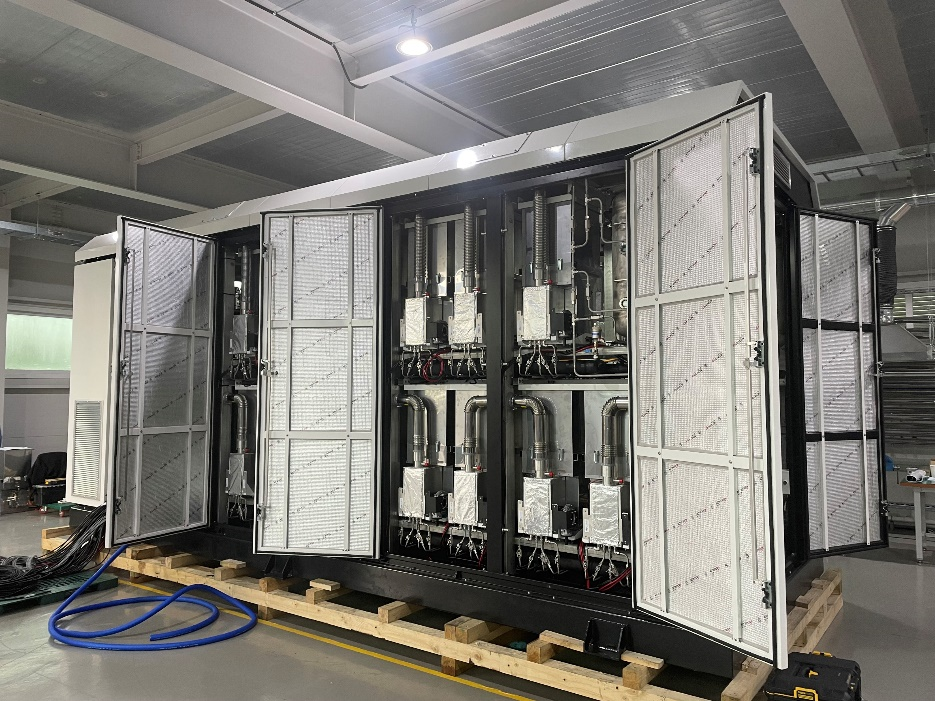
Gift of Fuel Cell Units Enhances UConn’s Clean Energy Commitment
Eight solid oxide fuel cell units will be donated to the Center for Clean Energy Engineering
April 2, 2024 | Matt Engelhardt